|
|
Hello, I hope everyone is enjoying the Spring season! Project highlight
Kristína Lidayová from the BIIF team developed a pipeline to align correlative light and electron microscopy (CLEM) images for ultrastructural mapping of protein distributions in cells. It aligns high-resolution TEM images with fluorescence microscopy channels using fiducial markers. Light microscopy (LM) provides multi-channel protein labeling, while electron microscopy (EM) offers nanoscale morphological details. Aligning these images enables comprehensive insights into cellular structures. Key aspects of the image analysis pipeline include:
Explore the code here If you have similar microscopy images or any image analysis questions feel free to reach us at biif@scilifelab.se or submit your project requests at https://nbis.se/services/bioimage-informatics. See more of our work on the BIIF projects page. Publication highlight
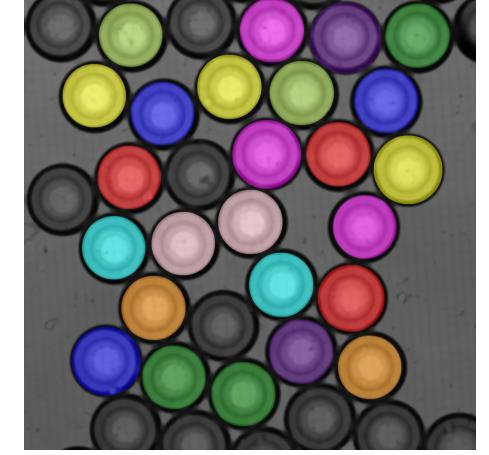
Jonas Windhager from the BIIF team supported a study utilizing a novel droplet microfluidics method to detect rare resistant E. coli subpopulations in bloodstream infections, addressing the clinical challenge of antibiotic heteroresistance. He developed a custom computational pipeline in Python to automate the analysis of time-lapse microscopy. By leveraging tunable image segmentation algorithms to quantify droplet shrinkage, the method successfully identified resistant cells at frequencies as low as 10-6, demonstrating a robust, high-throughput approach for improved diagnostic sensitivity. Interested in the image analysis pipeline? Check out the code More new findings from this project are coming soon — stay tuned for updates! Pre-Focus on Microscopy (FOM) discussionWe were delighted to host Dr. Oliver Biehlmaier and Dr. Alexia Loynton-Ferrand from the University of Basel. The visit included fruitful discussions on image analysis infrastructure, smart microscopy, and the tools used at our respective facilities. Upcoming eventsGloBIAS seminarAs part of the Global BioImage Analysts' Society (GloBIAS) seminar series, Nicholas Condon from the Institute for Molecular Bioscience, UQ, Australia will be speaking about "Big data infrastructure strategies for image analysis in a core facility" on 1st April, 2026 at 13:00 CEST. Register here Image data community days 2026Euro-Bioimaging is organizing Image Data Community Days from 13-17th April, 2026 with a list of lectures, hands-on workshops and networking sessions focussed on image data. Don't miss Jonas Windhager giving an update on the next major release of TissUUmaps in his talk "TissUUmaps 4: Facilitating Community Contributions to Spatial Biology Visualization" on 15th April, 2026 at 09:40 - 10:05 CET. Register here MIMER AI factory plenary kickoffInterested in using AI for research? Curious about how AI is being applied in life sciences and other domains? This event might be for you. Register by 15th April, 2026 19th OME Community MeetingOpen Microscopy Environment (OME) Community Meeting will be held at Düsseldorf from 28-30th April, 2026. More information here. NBIS workshop in Neural Nets and Deep LearningChristophe Avenel from the BIIF team along with other NBIS members is organizing an in-person course on "Neural Nets and Deep Learning" at Linköping University from 4-8th May, 2026. Check out the link for more details. Apply by 4th April, 2026 NBIS workshopNBIS training is offering the following courses in April and May:
For more details on training please check this page. NBIS drop-insA weekly drop-in that is open to any researcher with questions not only on image analysis but on bioinformatics, data management or HPC. When: Tuesdays at 14:00 CEST Where: https://meet.nbis.se/dropin to join the drop-in online There are also options to attend on-site in Stockholm, Lund, Chalmers and Gothenburg. For more details check out this page. Regards, Suganya Sivagurunathan On behalf of the BIIF team |
Newsletter history
-
April 2026
-
March 2026
-
February 2025
-
January 2025
-
October 2024
-
September 2024
-
June 2024
-
May 2024
-
February 2024
-
January 2024
-
December 2023
-
December 2023
-
November 2023
-
October 2023
-
August 2023
-
June 2023
-
February 2023
-
January 2023
-
December 2022
-
December 2022
-
October 2022
-
September 2022
-
May 2022
-
April 2022
-
March 2022
-
February 2022
-
January 2022
-
December 2021
-
November 2021
-
October 2021
-
September 2021
-
August 2021
-
June 2021
-
May 2021
-
April 2021
-
March 2021
-
February 2021
-
January 2021