Illustration (click to hide):
Project Description
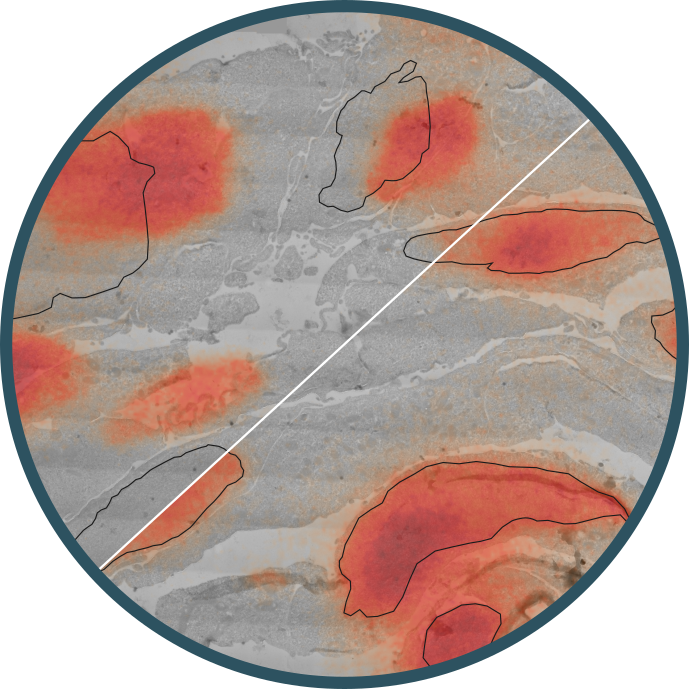
The project developed a correlative light and electron microscopy (CLEM) workflow for ultrastructural mapping of protein distributions in cells. It aligns high-resolution TEM images with fluorescence microscopy channels using fiducial markers. LM provides multi-channel protein labeling through fluorescence microscopy, while EM offers nanoscale morphological details. Aligning these images enables comprehensive insights into cellular structures.
Key aspects of the image analysis pipeline include:
• Fiducial particles detection in EM images using the template matching algorithm
• Fiducial particles detection in LM images using the Big-FISH Python package
• Finding correlation between EM and LM images using the Coherent Point Drift (CPD) registration
Project Information
-
BIIF Principal Investigators
- Kristína Lidayová
External Authors
Jan van der Beek -
Date
2024-07-02 🠚 2025-07-17 - GitHub page